Software
We open-sourced a variety of software packages. Many of them focusing on image registration. See the detailed descriptions as tiles and quick links to the respective GitHub repositories below.
Quick links
| Software | Description |
|---|---|
| SimpleClick | Interactive image segmentation |
| Aladdin | Deep-learning based atlas building |
| Easyreg | Deep-learning image registration toolbox |
| ICON | Deep-learning based inverse consistent image registration |
| LTS | Local temperature scaling |
| OAI Analysis 2 | Image analysis tools for the Osteoarthritis Initiative image dataset |
| Pediatric Airway Atlas | Software tools to build and query an atlas of the upper airway in children |
| RobOT | Point cloud registration |
| VoteNet | A deep learning label fusion approach for multi-atlas segmentation |
| YETI | A deep learning approach for perfusion imaging |
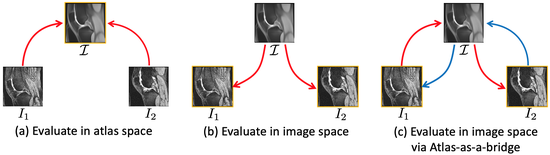
Aladdin
This software allows for joint AtLAs builDing and Diffeomorphic regIstration learNing (Aladdin) with pairwise alignment. In contrast to existing atlas-building approaches it uses the atlas as a bridge and incorporates pairwise similarity measures between images which are related indirectly through their atlas registrations.
ICON
ICON (Inverse COnsistent RegistratioN) is a non-parametric deep learning registration approach which relies only on inverse consistency for regularity. As the regularization neither involves explicit smoothing or a penality on spatial derivatives no affine pre-registration is required.
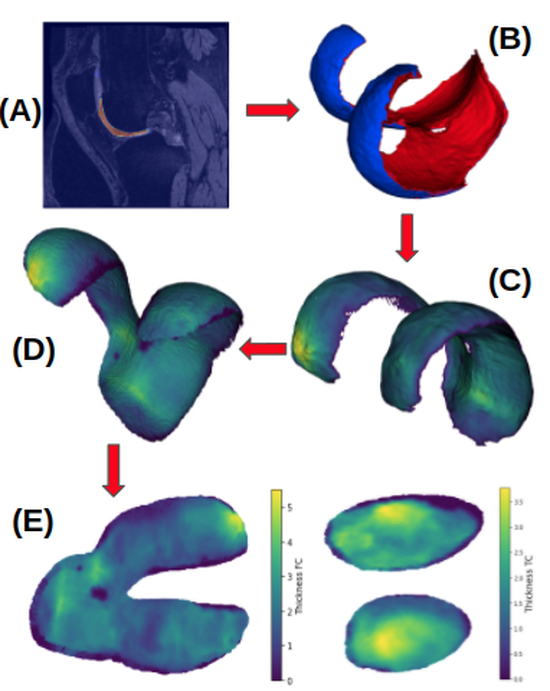
OAI Analysis 2
This software contains open-source analysis approaches for the Osteoarthritis Initiative (OAI) magnetic resonance image (MRI) data. The analysis code is largely written in Python with the help of ITK and VTK for data I/O and mesh processing as well as PyTorch for the deep learning approaches for segmentation and registration.
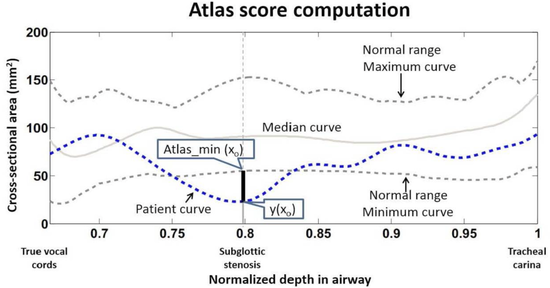
Pediatric Airway Atlas
The project aims to provide an analytical for measuring the normality of children’s airways. We build an age-based atlas on multiple CT images of normal subjects. First, we use a segmentation model to extract the airway.
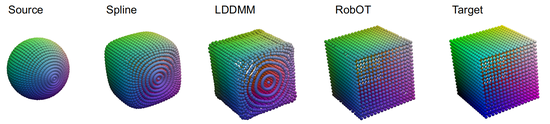
RobOT
This software provides a general framework for point cloud/mesh registration based on robust optimal mass transport (robOT) / unbalanced optimal mass transport. It supports both optimization- and learning-based registration approaches. It also provides a general framework for deep prediction tasks, e.
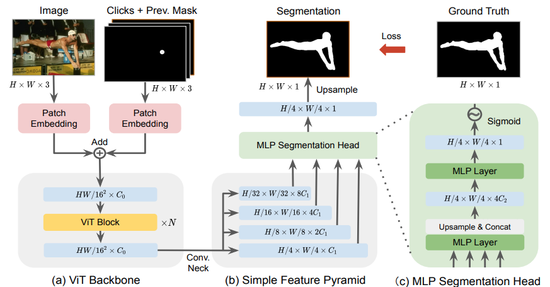
SimpleClick
This software allows for interactive image segmentation. We aim to develop SimpleClick as a practical tool for interactive image segmentation, editing, and generation. The SimpleClick repository can be found here: https://github.
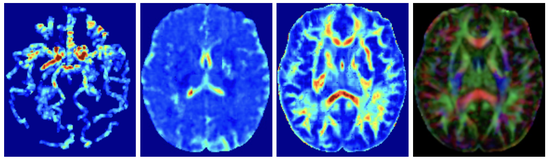
YETI
This software uses a learning framework (YETI) building on an auto-encoder structure between 2D and 3D image time-series, which incorporates an advection-diffusion model to capture blood perfusion. To help with identifiability, the deep learning model is trained via simulated data from an advection-diffusion simulator.
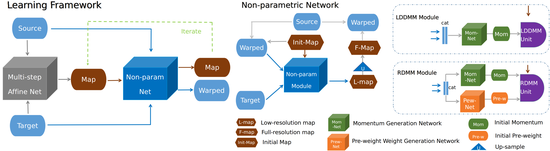
easyreg
EasyReg is an extension that builds on Mermaid, providing a simple interface to Mermaid and other popluar registration packages. The currently supported methods include Mermaid-optimization (i.e., optimization-based registration) and Mermaid-network (i.
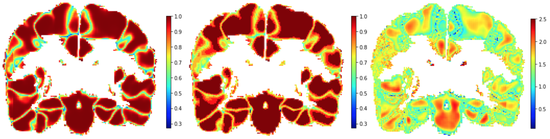
Local Temperature Scaling
Deep segmentation networks typically create their outputs via a softmax. Hence, the assignment appears to be probabilistic. However, it has been shown in literatute that interpreting these softmax outputs as label probabilities is not reliable and that they tend to be overly confident.

Mermaid
Mermaid: iMagE Registration via autoMAtIc Differentiation Mermaid is a registration toolkit making use of automatic differentiation for rapid prototyping. It includes various image registration models. In particular, stationary velocity field models (both based on velocity fields and momentum fields), scalar vector momentum Large Displacement Diffeomorphic Metric Mapping (LDDMM) models as well as the more generalized Region-specific Diffeomorphic Metric Mapping model (RDMM).
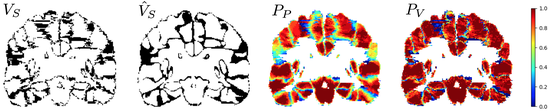
VoteNet
VoteNet is a deep-learning-based label fusion strategy for multi-atlas segmentation (MAS) which locally selects a set of reliable atlases whose labels are then fused via plurality voting. By selecting a good initial atlas set MAS with VoteNet significantly outperforms a number of other label fusion strategies as well as a direct deep-learning (DL) segmentation approach.